Replication-guided nucleosome packing and nucleosome breathing expedite the formation of dense arrays
The condensation of eukaryotic chromatin into DNA entails the formation of dense nucleosome arrays. These are frequently disrupted by transcription and replication, such that reassembly is required. The kinetics of this reassembly is of central interest. We investigate scenarios that enable to reach densely packed arrays within biologially reasonable timescales and find that nucleosome breathing, stepwise nucleosome assembly as well as replication guided processes expedite the array assembly.
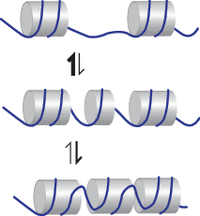
In a simplified way nucleosomes on DNA can be modeled as 1D particles, each occupying 147 base pairs. The assembly kinetics of such hard, mutually exclusive particles exhibit a 'jamming effect' that prevents reaching tightly packed arrays quickly. However, a more faithful description of nucleosomes takes into account that the outer parts of DNA can transiently unwrap (breathing) and that nucleosome assembly occurs in a stepwise manner. Both effectively makes nucleosomes soft particles. We show that, besides describing steady state patterns more accurately, softness expedites the array assembly kinetics considerably, because nucleosomes can squeeze into smaller gaps and make use of space that remains otherwise unoccupied.
We also discuss scenarios how the progression of the replication fork can promote rapid reassembly in its wake. For example, tight packing arises naturally if the fork progresses slowly compared to the reassembly rate.
B. Osberg, J. Nuebler, P. Korber and U. Gerland (2014),
Replication-guided nucleosome packing and nucleosome breathing expedite the formation of dense arrays,
Nucleic Acids Research 42 , 13633-13645.