Experimental Laboratory
MetA is a “thermal fuse” that inhibits growth and protects Escherichia coli at elevated temperatures

Adaptive stress resistance in microbes is mostly attributed to the expression of stress response genes, including heat-shock proteins. Here, we report a response of E. coli to heat stress caused by degradation of an enzyme in the methionine biosynthesis pathway (MetA). While MetA degradation can inhibit growth, which by itself is detrimental for fitness, we show that it directly benefits survival at temperatures exceeding 50°C, increasing survival chances by more than 1,000-fold. Using both experiments and mathematical modeling, we show quantitatively how protein expression, degradation rates, and environmental stressors cause long-term growth inhibition in otherwise habitable conditions. Because growth inhibition can be abolished with simple mutations, namely point mutations of MetA and protease knockouts, we interpret the breakdown of methionine synthesis as a system that has evolved to halt growth at high temperatures, analogous to “thermal fuses” in engineering that shut off electricity to prevent overheating.
Slower growth of Escherichia coli leads to longer survival in carbon starvation due to a decrease in the maintenance rate
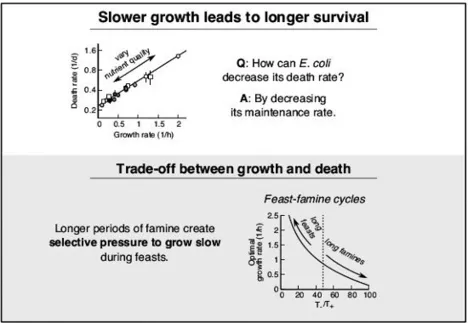
Fitness of bacteria is determined both by how fast cells grow when nutrients are abundant and by how well they survive when conditions worsen. Here, we study how prior growth conditions affect the death rate of Escherichia coli during carbon starvation. We control the growth rate prior to starvation either via the carbon source or via a carbon‐limited chemostat. We find a consistent dependence where death rate depends on the prior growth conditions only via the growth rate, with slower growth leading to exponentially slower death. Breaking down the observed death rate into two factors, maintenance rate and recycling yield, reveals that slower growing cells display a decreased maintenance rate per cell volume during starvation, thereby decreasing their death rate. In contrast, the ability to scavenge nutrients from carcasses of dead cells (recycling yield) remains constant. Our results suggest a physiological trade‐off between rapid proliferation and long survival. We explore the implications of this trade‐off within a mathematical model, which can rationalize the observation that bacteria outside of lab environments are not optimized for fast growth.
Death Rate of E. coli during Starvation Is Set by Maintenance Cost and Biomass Recycling
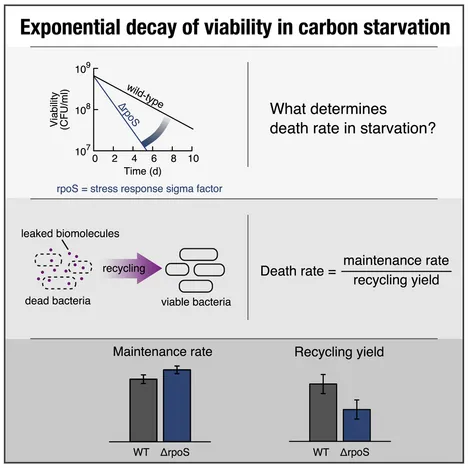
To break down organismal fitness into molecular contributions, costs and benefits of cellular components must be analyzed in all phases of the organism’s life cycle. Here, we establish the required quantitative approach for the death phase of the model bacterium Escherichia coli. We show that in carbon starvation, an exponential decay of viability emerges as a collective phenomenon, with viable cells recycling nutrients from cell carcasses to maintain viability. The observed collective death rate is determined by the maintenance rate of viable cells and the amount of nutrients recovered from dead cells. Using this relation, we study the cost of a wasteful enzyme during starvation and the benefit of the stress response sigma factor RpoS. While the enzyme increases maintenance and thereby the death rate, RpoS improves biomass recycling, decreasing the death rate. Our approach thus enables quantitative analyses of how cellular components affect the survival of non-growing cells.
A global resource allocation strategy governs growth transition kinetics of Escherichia coli
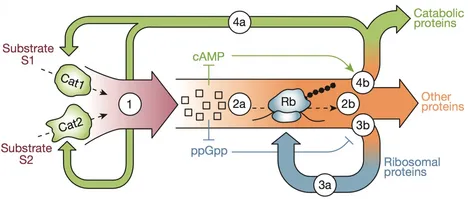
A grand challenge of systems biology is to predict the kinetic responses of living systems to perturbations starting from the underlying molecular interactions. Changes in the nutrient environment have long been used to study regulation and adaptation phenomena in microorganisms and they remain a topic of active investigation. Although much is known about the molecular interactions that govern the regulation of key metabolic processes in response to applied perturbations, they are insufficiently quantified for predictive bottom-up modelling. Here we develop a top-down approach, expanding the recently established coarse-grained proteome allocation models from steady-state growth into the kinetic regime. Using only qualitative knowledge of the underlying regulatory processes and imposing the condition of flux balance, we derive a quantitative model of bacterial growth transitions that is independent of inaccessible kinetic parameters. The resulting flux-controlled regulation model accurately predicts the time course of gene expression and biomass accumulation in response to carbon upshifts and downshifts (for example, diauxic shifts) without adjustable parameters. As predicted by the model and validated by quantitative proteomics, cells exhibit suboptimal recovery kinetics in response to nutrient shifts owing to a rigid strategy of protein synthesis allocation, which is not directed towards alleviating specific metabolic bottlenecks. Our approach does not rely on kinetic parameters, and therefore points to a theoretical framework for describing a broad range of such kinetic processes without detailed knowledge of the underlying biochemical reactions.