New Publication in the Journal of Proteome Research
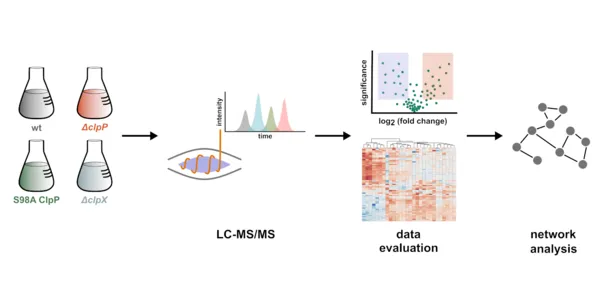
Staphylococcus aureus represents an opportunistic pathogen, which utilizes elaborate quorum sensing mechanisms to precisely control the expression and secretion of virulence factors. Previous studies indicated a role of the ClpXP proteolytic system in controlling pathogenesis. While detailed transcriptome data for S. aureus ClpP and ClpX knockout mutants is available, corresponding studies on the proteome and secretome level are largely lacking. To globally decipher the functional roles of ClpP and ClpX, we utilized S. aureus genomic deletion mutants of the corresponding genes for in-depth proteomic liquid chromatography–mass spectrometry (LC–MS)/MS analysis. These studies were complemented by an inactive ClpP active-site mutant strain to monitor changes solely depending on the activity and not the presence of the protein. A comparison of these strains with the wildtype revealed, e.g., downregulation of virulence, purine/pyrimidine biosynthesis, iron uptake, and stress response. Correspondingly, the integration of metabolomics data showed a reduction in the subset of purine and pyrimidine metabolite levels. Interestingly, a comparison between the ClpP knockout and ClpP S98A active-site mutant strains revealed characteristic differences. These results are not only of fundamental importance to understand the cellular role of ClpXP but also have implications for the development of novel virulence inhibitor classes.
Kirsch, V., Fetzer, C., Sieber, S. A., "A global inventory of ClpP and ClpX regulated proteins in Staphylococcus aureus", J. Proteome Res.