Chemical proteome mining

The Challenge:
Up to 50% of cellular proteins still lack an assigned functional role. Given that these proteins of unknown function play crucial roles in cellular physiology, deciphering their exact function together with the design of tailored inhibitors would increase our knowledge and repertoire of druggable targets.
Our Goals:
We aim to design small molecule probes and apply them in the mining of complex proteomes for proteins bearing conserved chemical features. Here, especially cofactor-dependent enzymes stick out due to their essential role in many cellular pathways. A chemical tool reporting on such an enzyme class would reveal proteins with known as well as unknown cellular roles. Special emphasis is given to those proteins that are uncharacterized, but have putative roles in disease, revealing novel targets for therapeutic intervention.
Our Research:
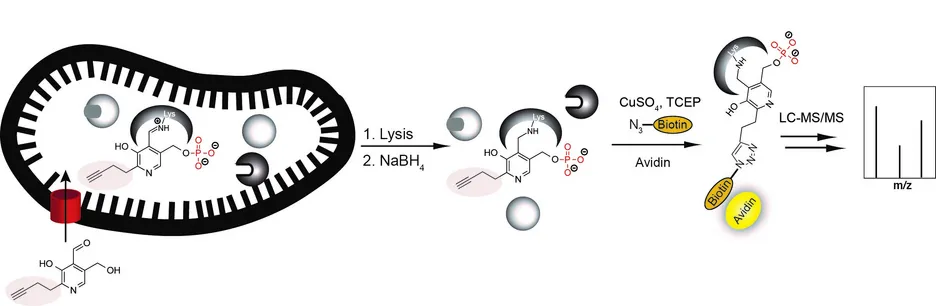
Based on the cofactor pyridoxal phosphate (PLP), we synthesized tailored probes which infiltrate the cellular metabolic machinery of bacterial and human cells. The probes get phosphorylated within the living cell and are subsequently incorporated in PLP-dependent enzymes. Enrichment and identification of labeled proteins via mass spectrometry revealed a wealth of known and unknown PLP-dependent targets with essential cellular roles. Follow up studies focus on the design of specific inhibitors of this enzyme class as novel antibiotics. All our target identifications and full proteome analyses are carried out via gel-free proteome measurements on Orbitrap XL, Orbitrap Fusion and Q-Exactive instruments. Latest methods for quantification are implemented (isotopic labeling as well as label-free strategies) and provide us with full datasets that we upload for public access in the PRIDE database. In addition, we provide resources, e.g. to determine the cleavage specificity of proteases and the background labeling of photocrosslinkers in whole proteomes. These tools can be found under resource.
Selected publication:
1. Hoegl, A., Nodwell, M.B., Kirsch, V.C., Bach, N.C., Pfanzelt, M., Stahl, M., Schneider, S., and Sieber, S.A. "Mining the cellular inventory of pyridoxal phosphate-dependent enzymes with functionalized cofactor mimics." Nat. Chem. 2018, 10, 1234-1245. PMID: 30297752, PMCID: PMC6252082
2. Kielkowski, P., Buchsbaum, I.Y., Kirsch, V.C., Bach, N.C., Drukker, M, Cappello, S.*, Sieber, S.A*, "FICD activity and AMPylation remodelling modulate human neurogenesis", Nat. Commun., 2019. PMID: 31980631, PMCID: PMC6981130
3. Fux, A., Sieber, S.A., "Biochemical and proteomic studies of human pyridoxal 5’-phosphate-binding protein (PLPBP)", ACS Chem. Biol., 2020, 15, 254-261. PMID: 31825581
4. Kielkowski, P., Buchsbaum, I.Y., Becker, T., Bach, K., Cappello, S., Sieber, S.A., "A pronucleotide probe for live cell imaging of protein AMPylation", ChemBioChem, 2020, 21(9), 1285-1287. PMID: 32027064