Jobs
For postdoctoral researchers
Research Associate/Postdoc in Computational Biophysics Technical University of Munich
A Research Associate position is available in computational biophysics at the Center of Functional Protein Assemblies of TU Munich, Germany, to start immediately. The position is intended to offer a long term perspective (initially for 3 years but extendable for up to 5 years). It includes the possibility to participate in supervision of PhD and master students to develop an independent research profile and to possibly apply for third party funding. It also involves participation in teaching within the physics/biophysics curriculum.
The research focus shall be in the area of studying biomolecular structure and dynamics employing molecular mechanics or mixed quantum/classical mechanics. In addition, it involves machine learning and artificial intelligence approaches for force field design and prediction of biomolecular binding. Possible research directions can focus on better understanding of protein-protein, protein-membrane and/or protein-DNA/RNA interactions using advanced sampling methods and free energy calculations. It also includes computational design of new protein-protein interactions, new protein based materials and drug design in close collaboration with experimental groups on the TU Munich campus.
Successful candidates should have a PhD in either physics, physical chemistry, biophysics or computational science. Experience in the area of biomolecular simulations, QM/MM methods and/or machine learning approaches is a plus. The ability to handle multiple projects simultaneously is also desired.
The Technical University of Munich belongs to the scientific top addresses and is one of the Universities of Excellence in the Federal Republic of Germany. The Zacharias group closely collaborates with experimental groups to better understand the dynamics and association of biomolecules. State-of-the-art computer equipment is available including access to supercomputing facilities. The position will be paid according to the German E13 level.
Please, send your CV and cover letter describing your research interests including the addresses of two referees to (preferably by e-mail):
Prof. Dr. Martin Zacharias
Chair of Theoretical Biophysics-Molecular Dynamics
Center of Functional Protein Assemblies
Technical University of Munich
Ernst-Otto-Fischer-Str. 8
D-85748 Garching, Germany
e-mail: Zacharias@tum.de
For doctoral researches
For B.Sc. and M.Sc. Students
2. Modellierung der Proteinfaltung mit vergröberten Proteinmodellen
Please switch to German language
1. Structure prediction of Protein assemblies based on chemical crosslinking
Most cellular processeand are mediated by large protein complexes (assemblies) and are influenced by transient protein-protein interactions. By using chemical crosslinking it is possible to trapp and analyse these interactions in vitro (in a test tube) but also in vivo (in the cell). Cross linking refers to the covalent connection of two proteins if they come close together (to a distance of less than ~2-3 nm). The crosslink positions on the protein surface can be identified by mass spectrometry. With the knowledge of a sufficient number of such cross links it is in principle possible to determine how the proteins interact and to create a model of the spatial arrangements of the proteins in the molecular assembly or complex. Goal of the master thesis is to design and improve approaches to predict the structure of multi protein complexes based on the structure of the individual proteins and based on the crosslinking data. In the master thesis computational approaches will be used and some knowledge of a programming language like Python is an advantage. If successful the work can have great impact on better understanding biophysical processes in a cell.
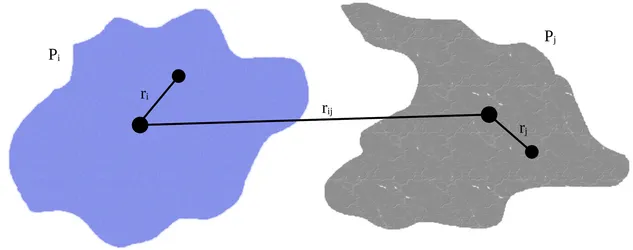
2. Describing Proteins as Anisotropic Coarse Densities
Protein-protein interactions play a central role in life, but understanding these interactions in crowded environments and at larger scales is challenging in simulations. In the last decades, considerable effort has been devoted to the development of coarse-grained force fields that describe proteins and other cellular components with large isotropically interacting beads to simplify the simulation systems and speed up computation. However, the loss of directionality, charge distribution, and deformation due to conformational changes in this coarse description is often problematic in practice and hinders the scientific output of these methods. For this reason, we have recently developed a new approach for coarse-grained simulations that describes proteins as arbitrarily shaped particle densities interacting with each other. This master’s thesis involves the testing and benchmarking of this new method, as well as contributing to the development of simulation and visualization software in Python.