Wolfgang Heinemeyer
Since completing my doctorate, I have been working with the yeast Saccharomyces cerevisiae as a model organism in the field of essential cellular regulation by the ubiquitin/proteasome system. In eukaryotic cells, this system fulfils a multitude of vital functions, ranging from protein quality control to regulation of the cell cycle and, in vertebrates, antigen presentation. It is now known that disturbances in this system lead to serious human diseases such as cancer, neurodegeneration or immune defects.
Selective labelling by polyubiquitin chains directs proteins for degradation to oligopeptides by the 26S proteasome. This high molecular weight protease consists of the cylindrical 20S core particle (CP), which hosts proteolytic activities, and 19S regulatory units, which are responsible for the recognition of polyubiquitinated substrates, their unfolding and translocation into the 20S complex.
At the beginning of my work in the early 1990s, I was able to prove the general function of the proteasome in ubiquitin-mediated proteolysis in vivo for the first time using mutants with defects in one of the three proteolytic activities of the CP. Such mutants then also allowed the identification of the first, defined in-vivo substrates of the proteasome.
The genes encoding the affected subunits in the mutants were cloned and shown to be essential for yeast viability. All cloned 20S proteasome genes turned out to constitute a gene family of α-type and β-type members, the number of which I was able to complete to 14 by cloning two further yeast α-type proteasome genes. The structural model for the CP derived by us proved to be true as a four-layered cylinder with twice 14 different subunits, of which 7 α-type subunits occupy fixed positions in the outer ring structures and 7 β-type subunits form the two inner rings.
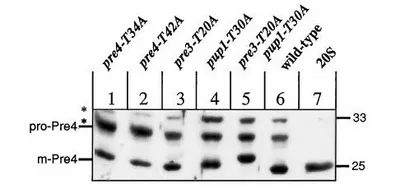
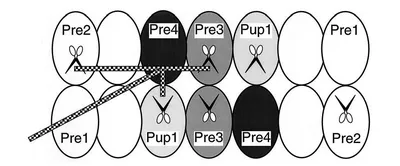
Through site-directed mutagenesis, three of the 7 β-type subunits were classified as N-terminal threonine proteases. With the help of short model peptide substrates, different specificities could be assigned to the three proteolytic centers, which were characterised in preferential cleavage of peptide bonds C-terminal to hydrophobic, basic or acidic amino acids. As part of a collaboration involving Michael Groll and Robert Huber, the qualitative analysis of the cleavage products generated in vitro by wild-type and mutant proteasomes from a large protein allowed a detailed determination of sequence motifs that determine cleavage at the different centers. Furthermore, the mechanism of self-activation of active β-type subunits, which involves the cleavage of N-terminal propeptides, could be elucidated using X-ray structure analysis of purified mutant proteasomes. The participation of activated subunits in the maturation of non-active β-type subunits, which I had postulated, was also confirmed in this collaboration. Furthermore, we identified different contributions of the propeptides of active subunits in proteasome assembly as well as a common function of the propeptides, namely the prevention of a modification at the N-terminal threonine, which would lead to loss of activity.
After joining the group of Michael Groll at the TUM, I continued research on the CP by generating mutants of the yeast particle, which in part were also suited to mimic structural elements found in the constitutive or immuno-proteasome of vertebrates. Purified proteasome from such mutants was used for x-ray crystallographic analysis to address several questions concerning i) the general mechanism of substrate proteolysis and autocatalytic propeptide removal, ii) the binding mode and subunit specificity of different proteasome inhibitors, iii) the structural basis for resistance to the clinically applied inhibitor bortezomib in human cell lines or iiii) the molecular explanation for severe immunodeficiency found in a mouse mutant and the severe auto-inflammantory syndrome in a human hereditary disease.
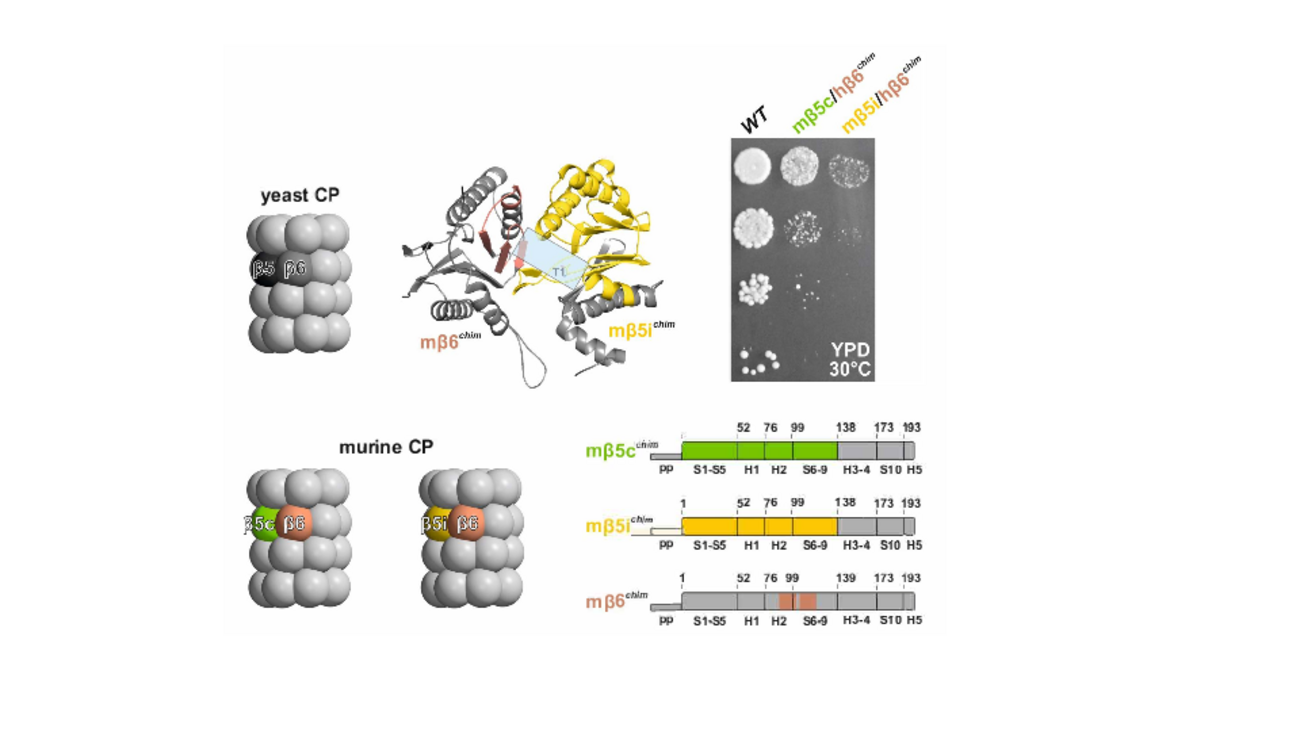
A large collection of yeast 20S proteasome mutants expressing chimeric yeast/mammalian subunits was generated to elucidate not only the reason for selectivity of inhibitors to either the mammalian constitutive or immuno-proteasome but also to successfully narrow down the sequence requirements leading to the widened substrate binding pocket of the vertebrate β5i immuno-proteasome subunit compared to the more closed pocket in β5 of constitutive proteasomes.
In addition, I participated in various projects of group members by constructing yeast strains expressing tagged versions of proteins under study or by providing systems for overexpression of heterologous proteins in yeast.
Publications
Xin B. T., Huber E. M., de Bruin G., Heinemeyer W., Maurits E., Espinal C., Du Y., Janssens M., Weyburne E. S., Kisselev A. F., Florea B. I., Driessen C., van der Marel G. A., Groll M. and Overkleeft H. S.
Structure-Based Design of Inhibitors Selective for Human Proteasome β2c or β2i Subunits
J. Med. Chem., 2019, 62, 1626-1642, PDF
Treise I., Huber E. M., Klein-Rodewald T., Heinemeyer W., Grassmann S. A., Basler M., Adler T., Rathkolb B., Helming L., Andres C., Klaften M., Landbrecht C., Wieland T., Strom T. M., McCoy K. D., Macpherson A. J., Wolf E., Groettrup M., Ollert M., Neff F., Gailus-Durner V., Fuchs H., Hrabě de Angelis M., Groll M. and Busch D. H.
Defective immuno- and thymoproteasome assembly causes severe immunodeficiency
Sci. Rep., 2018, 8, (5975), 1-18, PDF
Cui H., Baur R., Le Chapelain C., Dubiella C., Heinemeyer W., Huber E. M. and Groll M.
Structural Elucidation of a Nonpeptidic Inhibitor Specific for the Human Immunoproteasome
Chembiochem., 2017, 18, 523-526, PDF
Huber E. M., Heinemeyer W., de Bruin G., Overkleeft H. S. and Groll M.
A humanized yeast proteasome identifies unique binding modes of inhibitors for the immunosubunit β5i
EMBO J., 2016, 35, 2602-2613, PDF
Huber E.M., Heinemeyer W., Li X., Arendt C. S., Hochstrasser M. and Groll M.
A unified mechanism for proteolysis and autocatalytic activation in the 20S proteasome
Nat. Commun., 2016, 7, 1-10, PDF
Huber E. M., de Bruin G., Heinemeyer W., Paniagua Soriano G., Overkleeft H. S. and Groll M.
Systematic Analyses of Substrate Preferences of 20S Proteasomes Using Peptidic Epoxyketone Inhibitors
J. Am. Chem. Soc., 2015, 137, 7835-7842, PDF
Huber E. M., Heinemeyer W. and Groll M.
Bortezomib-resistant mutant proteasomes: structural and biochemical evaluation with carfilzomib and ONX 0914
Structure, 2015, 23, 407-417, PDF
Beck P., Heinemeyer W., Späth A. L., Elnakady Y., Müller R. and Groll M.
Interactions of the natural product kendomycin and the 20S proteasome
J. Mol. Biol., 2014, 426, 3108-3117, PDF
Huber E. M., Basler M., Schwab R., Heinemeyer W., Kirk C. J., Groettrup M. and Groll M.
Immuno- and constitutive proteasome crystal structures reveal differences in substrate and inhibitor specificity,
Cell, 2012, 148, 727-738, PDF
Heinemeyer W., Ramos P. C. and Dohmen R. J.
The ultimate mincer at nano-scale: Assembly structure and active sites of the 20S proteasome core
Cell. Mol. Life Sci., 2004, 61, 1562-1578,
Jäger S., Strayle J., Heinemeyer W. and Wolf D. H.
Cic1, an adapter protein specifically linking the 26S proteasome to its substrate, the SCF component Cdc4
EMBO J., 2001, 20, 4423-4431,
Heinemeyer W.
The active sites and assembly of the 20S proteasome, in: "Proteasomes: The world of regulatory proteolysis"
W. Hilt and D. H. Wolf (eds.), Eurekah.com / Landes Bioscience, Austin / Georgetown, Texas, USA, pp 48-70, 2000
Groll M., Heinemeyer W., Jäger S., Ullrich T., Bochtler M., Wolf D. H. and Huber R.
The catalytic sites of 20S proteasomes and their role in subunit maturation: A mutational and crystallographic study
Proc. Natl. Acad. Sci. USA, 1999, 96, 10976-83, PDF
Jäger S., Groll M., Huber R., Wolf D. H. and Heinemeyer, W.
Proteasome beta-type subunits: Unequal roles in core particle maturation and a hierarchy of active site function
J. Mol. Biol., 1999, 291, 997-1013, PDF
Nussbaum A. K., Dick T. P., Keilholz W., Schirle M., Stevanovic S., Dietz K., Heinemeyer W., Groll M., Wolf D. H., Huber R., Rammensee H.-G. and Schild H.
Cleavage motifs of the yeast 20S proteasome beta subunits deduced from digests of enolase I
Proc. Natl. Acad. Sci. USA, 1998, 95, 12504-9, PDF
Dick T. P., Nussbaum A. K., Deeg M., Heinemeyer W., Groll M., Schirle M., Keilholz W., Stevanovic S., Wolf D. H., Huber R., Rammensee H.-G. and Schild H.
Contribution of proteasomal beta-subunits to the cleavage of peptide substrates analyzed with yeast mutants
J. Biol. Chem., 1998, 273, 25637-46, PDF
Ditzel L., Huber R., Mann K., Heinemeyer W., Wolf D. H. and Groll M.
Conformational constraints for protein self-cleavage in the proteasome
J. Mol. Biol., 1998, 279, 1187-1191, PDF
Heinemeyer W., Fischer M., Krimmer T., Stachon U. and Wolf D. H.
The active sites of the eukaryotic 20S proteasome and their involvement in subunit precursor processing
J. Biol. Chem., 1997, 272, 25200-25209, PDF
Hilt W., Heinemeyer W. and Wolf, D. H.
The proteasome and protein degradation in yeast
Adv. Exp. Med. Biol., 1996, 389, 197-202, PDF
Heinemeyer W., Tröndle N., Albrecht G. and Wolf D. H.
PRE5 and PRE6, the last missing genes encoding 20S proteasome subunits from yeast? Indication for a set of 14 different subunits in the eukaryotic proteasome core
Biochemistry, 1994, 33, 12229-12237, PDF
Heinemeyer W., Gruhler A., Möhrle V., Mahé Y. and Wolf D. H.
PRE2, highly homologous to the human MHC-linked RING10 gene, codes for a yeast proteasome subunit necessary for chymotryptic activity and degradation of ubiquitinated proteins
J. Biol. Chem., 1993, 268, 5115-5120, PDF
Hilt W., Heinemeyer W. and Wolf, D. H.
Studies on the yeast proteasome uncover its basic structural features and multiple in vivo functions
Enzyme&Protein, 1993, 47, 189-201, PDF
Richter-Ruoff B., Heinemeyer W. and Wolf D. H.
The proteasome/ multicatalytic-multifunctional proteinase: in vivo function in the ubiquitin-dependent N_end rule pathway of protein degradation in eukaryotes
FEBS Lett., 1992, 302, 192-196, PDF
Heinemeyer W., Simeon A., Hirsch H. H., Schiffer H. H., Teichert U. and Wolf D. H.
Lysosomal and non-lysosomal proteolysis in the eukaryotic cell: studies on yeast
Biochem. Soc. Trans., 1991, 19, 724-725, PDF
Heinemeyer W., Kleinschmidt J. A., Saidowsky J., Escher C. and Wolf D. H.
Proteinase yscE, the yeast proteasome/multicatalytic-multifunctional proteinase: mutants unravel its function in stress induced proteolysis and uncover its necessity for cell survival
EMBO J., 1991, 10, 555-562, PDF
Alt-Moerbe J., Heinemeyer W. and Schröder J.
The virD genes from the vir region of the Ti plasmid: T-region border dependent processing steps in different rec mutants of Escherichia coli
Gene, 1990, 96, 43-49, PDF
Heinemeyer W., Buchmann I., Tonge D. W., Windass J. D., Alt-Moerbe J., Weiler E. W., Botz T. and Schröder J.
Two Agrobacterium tumefaciens genes for cytokinin-biosynthesis: Ti plasmid-coded isopentenyltransferases adapted for function in prokaryotic or eukaryotic cells
Mol. Gen. Genet., 1987, 210, 156-164, PDF
Heinemeyer W., Alt J. and Herrmann R. G.
Nucleotide sequence of the clustered genes for apocytochrome b6 and subunit 4 of the cytochrome b/f complex in the spinach plastid chromosome
Current Genetics, 1984, 8, 543-549, PDF