Melissa Gräwert
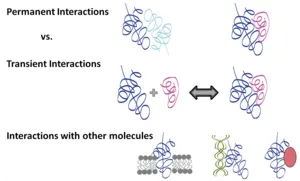
So far my major research interests have been the interactions between biological macromolecules that I have analyzed structurally and biophysically. Protein interactions are the fundamental driving force of almost every cellular process. Protein-Protein Interactions (PPIs) play an important role in cell cycle control, signal transduction and intermediary metabolism just to name a few. Thus, it is important to look beyond the function of every single gene product and to additionally understand how the ensemble of these gene products, subassemblies, and regulatory networks act together in highly evolved organisms. Macromolecular X-ray crystallography (MX) is a very important tool for these studies. This high resolution technique allows the visualization of interactions at an atomic level. From this, conclusions on binding affinities, forces and specificity can be derived. During my postdoctoral fellowship in Michael Groll’s lab, I was deeply involved in setting up and running our state-of-the-art X-ray core facility comprising an in-house X-ray generator and robotics for setting up crystallization trials. This infrastructure allowed us to grow many crystals and more importantly obtain high resolution structures that tell us fascinating stories. For example, co-crystallization of the 700 kDa proteasome from baker’s yeast with ligands resulted in important new structures with potential application as anti-cancer drugs. One of these gave insights into a new mode of proteasome inhibition by α-keto-aldehydes explaining their low levels of cross reactions and thus, reduced toxicity.
In successful collaborations with members of the Biophysics and Biotechnology department, I gained information on the driving forces of PPIs (visualization of oligomerization intermediates of a mutated Foldon protein, stability of PPIs (determination of quaternary contacts important for the specific polymerization of IgM which is responsible for its high overall avidity: insights on the structural evolution of the conserved antibody domains from shark with increased stability and secretion efficiency and specificity of PPIs (elucidation of the selective binding of Hsp90 and Hsp70 to different sites of the co-chaperone Sti1/Hop TPR domains).
Awards
2008 - 2012: Financial support by the Center for Integrated Protein Science Munich (26.000 €)
2008 - 2010: Postdoctoral stipend by the Peter und Traudl Engelhorn Foundation for “Exploiting nature's rich source of proteasome inhibitors as starting points in drug development”, Munich, Germany
Publications
Feige M. J., Gräwert M. A., Marcinowski M., Hennig J., Behnke J., Ausländer D., Herold E. M., Peschek J., Castro C. D., Flajnik M. F., Hendershot L. M., Sattler M., Groll M., Buchner J.
The structural analysis of shark IgNAR antibodies reveals evolutionary principles of immunoglobulins
Proc. Natl. Acad. Sci. USA, 2014, 111, 8155-60, PDF
Berthelmann A., Lach J., Gräwert M. A., Groll M., Eichler J.
Versatile C3-symmetric scaffolds and their use for covalent stabilization of the foldon trimer
Org. Biomol. Chem., 2014, 12, 2606-14, PDF
Müller R., Gräwert M. A., Kern T., Madl T., Peschek J., Sattler M., Groll M., Buchner J.
High-resolution structures of the IgM Fc domains reveal principles of its hexamer formation
Proc. Natl. Acad. Sci. USA, 2013, 110, 10183-8, PDF
Gräwert M. A., Groll M.
Eukaryotic 20S Proteasome
In: Handbook of Proteolytic Enzymes (Rawlings N. D. and Salvesen G. S., Eds.)
Oxford: Academic Press, 2013, 3684–91
Gräwert M. A., Groll M.
PhTET2 Aminopeptidase
In: Handbook of Proteolytic Enzymes (Rawlings N. D. and Salvesen G. S., Eds.)
Oxford: Academic Press, 2013, 1645–50
Schmid A. B., Lagleder S., Gräwert M. A., Röhl A., Hagn F., Wandinger S. K., Cox M. B., Demmer O., Richter K., Groll M., Kessler H., Buchner J.
The architecture of functional modules in the Hsp90 co-chaperone Sti1/Hop
EMBO J., 2012, 31, 1506-17, PDF
Gräwert M. A., Groll, M.
Exploiting nature's rich source of proteasome inhibitors as starting points in drug development
Chem. Commun., 2012, 48, 1364-78, PDF
Gräwert M. A., Gallastegui N., Stein M. L., Schmidt B., Kloetzel P.-M., Huber R., Groll M.
Elucidation of α-Keto-Aldehyde Binding Mechanism Reveals a Novel Lead Structure Motif for Proteasome Inhibition
Angew. Chem. Int. Ed., 2011, 50, 542-4 , PDF