Ingrid Span
Isoprenoids constitute the largest class of natural products with more than 55,000 known compounds, including molecules of vital importance such as cholesterol and carotenoids. In all three kingdoms of life, the universal precursors of isoprenoids are isopentenyl diphosphate (IPP) and dimethylallyl diphosphate (DMAPP). In mammals the mevalonate pathway is the unique source for these molecules. Plant chloroplasts, many pathogenic bacteria including Mycobacterium tuberculosis, and some protozoan, such as the malaria parasites Plasmodium falciparum, however, use an alternative pathway, 1-deoxy-D-xylulose 5-phosphate (DXP) pathway. This pathway is therefore an attractive target for herbicide, bactericide and anti-malarial drug development.
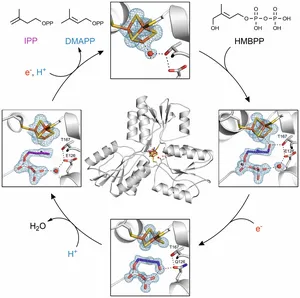
The final step of the DXP pathway is the conversion of (E)-1-hydroxy-2-methylbut-2-enyl 4-diphosphate (HMBPP) into a mixture of IPP and DMAPP. This reaction is catalyzed by the metalloenzyme IspH that incorporates an iron-sulfur cluster in the active site. This inorganic cofactor enables IspH to perform a sophisticated 2H+ /2e− reduction and deoxygenation reaction. When the substrate binds into the active site of the enzyme, a structural rearrangement occurs resulting in the closed protein conformation that protects the active site from the solvent. Initially, the substrate forms an alkoxide complex with the apical iron of the Fe4S4 cluster that is stabilized by threonine 167. Reduction of the cluster leads to rotation of the hydroxymethyl group, resulting in a cyclic conformation that is stabilized by several hydrogen bonds including one to glutamate 126. Electron transfer generates an allyl anion that is able to abstract a proton at two different positions affording a mixture of both products, IPP and DMAPP. This structural analysis, supported by spectroscopic data, have shed light on to the complex binding network between the substrate, the iron-sulfur cluster, the protein backbone and a water molecule in the active center of the enzyme.
These insights into the reaction mechanism on molecular level were obtained during my PhD thesis by combining protein crystallography with various spectroscopic techniques. With this information we started collaborating with Prof. Eric Oldfield from the University of Illinois at Urbana-Champaign with the goal of identifying and analyzing potent inhibitors for IspH and their structure-function relationship. Prof. Oldfield’s group synthesized various classes of molecules some of which had an effect on the activity of IspH. Within this project we learned many things about the binding and interactions of different functional groups with the iron-sulfur cofactor but also discovered an intriguing acetylene hydratase activity. With this work we hope to contribute to the development of drugs against infective diseases.
Awards
2013: Erlenmeyer prize for her Ph.D. thesis on: "Structural and functional characterisation of the iron-sulfur protein IspH in complex with ligands", Munich, Germany (1,500 €)
2012: Graduation prize Bund der Freunde der TUM for her Ph.D. thesis on “Structural and functional characterisation of the iron-sulfur protein IspH in complex with ligands”, Munich, Germany (1,500 €)
Publications
Span I., Wang K., Eisenreich W., Bacher A., Zhang Y, Oldfield E., Groll M.
Insights into the Binding of Pyridines to the Iron-Sulfur Enzyme ISPH
J. Am Chem Soc., 2014, 22, 7926-32, PDF
Span I., Wang K., Wang W., Jauch J., Eisenreich W., Bacher A., Oldfield E., Groll M.
Structures of fluoro, amino, and thiol inhibitors bound to the [Fe4S4] protein IspH
Angew. Chem. Int. Ed., 2013, 52, 2118-21, PDF
Span I., Wang K., Wang W., Zhang Y., Bacher A., Eisenreich W., Li K., Schulz C., Oldfield E., Groll M.
Discovery of Acetylene Hydratase Activity of the Iron-Sulfur Protein IspH
Nat. Commun., 2012, 3 (1042), 1-8, PDF
Wang W., Wang K., Span I., Jauch J., Bacher A., Groll M., Oldfield E.
Are Free Radicals Involved in IspH Catalysis? An EPR and Crystallographic Investigation
J. Am. Chem. Soc., 2012, 134, 11225-34, PDF
Span I., Gräwert T., Bacher A., Eisenreich W., Groll M.
Crystal structures of mutant IspH proteins reveal a rotation of the substrate's hydroxymethyl group during catalysis
J. Mol. Biol., 2012, 416, 1-9, PDF
Gräwert T., Span I., Bacher A., Groll M.
Reductive dehydroxylation of allyl alcohols by IspH protein
Angew. Chem. Int. Ed. Engl., 2010, 49, 8802-9, PDF
Gräwert T., Span I., Eisenreich W., Rohdich F., Eppinger J., Bacher A., Groll M.
Probing the reaction mechanism of IspH protein by x-ray structure analysis
Proc. Natl. Acad. Sci. USA, 2010, 107, 1077-81, PDF
Gräwert T., Rohdich F., Span I., Bacher A., Eisenreich W., Eppinger J., Groll M
Structure of active IspH enzyme provides mechanistic insights into substrate reduction
Angew. Chem. Int. Ed., 2009, 48, 5756-9, PDF