Research Overview
Below you can find a description of key concepts in our research. If you are interested in specific third-party funded projects we are currently working on, please follow this link to Current Projects.
In situ and Ambient Mass Spectrometry
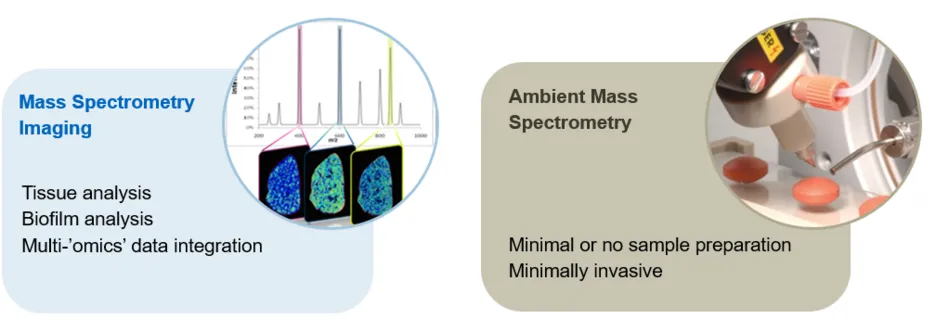
We develop in situ and ambient mass spectrometry methods to extend their range of application.
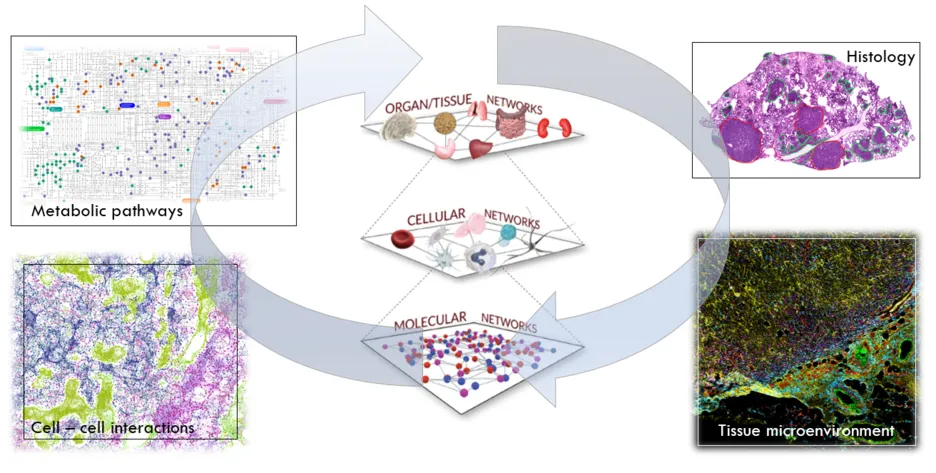
In situ mass spectrometry refers to techniques that analyse metabolites in their sites of origin, such as bacterial cultures on a Petri dish or endogenous metabolites in the tissue microenvironment. One of the main techniques we use for this purpose is imaging mass spectrometry (MSI). For MSI, samples are processed into thin sections using a cryostat, and then scanned with the ion source along a raster grid, recording a mass spectrum at each point. The grid points correspond to the pixels in the generated two-dimensional representation of the sample. We are currently using mainly Desorption Electrospray Ionisation (DESI) MSI and Matrix-assisted Laser Desorption Ionisation (MALDI) MSI to analyse endogenous metabolites and lipids. In addition, we develop workflows for multi-modal imaging and data integration in order to allow analysis of these multi-layered data to investigate complex relationships of metabolism with information such as morphology, tissue composition, or cell types.
Ambient mass spectrometry refers to MS techniques that analyse samples in their native form and under atmospheric conditions without any or only minimal sample preparation. We have extensive experience with DESI (the pioneer of ambient mass spectrometry) as well as Rapid Evaporative Ionization Mass Spectrometry (REIMS) and its use in profiling (fingerprinting) and screening applications such as screening nanoscaled reactions, microbial metabolites, or cell assays. Our newest acquisition is a Plasmion SICRIT dielectric barrier discharge ionisation source which allows direct analysis of the volatile sample fraction.
Experimental Toolbox
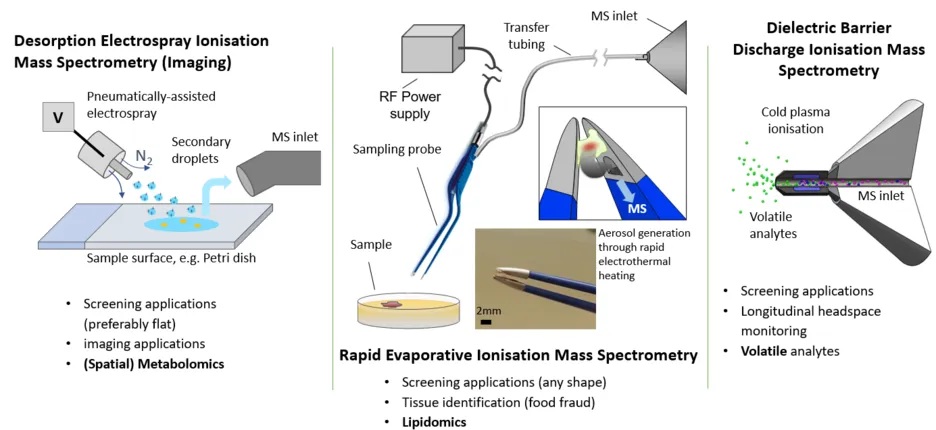
Our experimental toolbox consists of Desorption Electrospray Ionisation Mass Spectrometry (Imaging), Rapid Evaporative Ionisation Mass Spectrometry (REIMS) and the Plasmion SICRIT cold-plasma based ionisation source. Currently we are using a Thermo Q-Exactive Plus, a Thermo Exactive Classic and a Thermo LTQ XL ion trap for mass analysis. We further have an Agilent QTof 6224 for future hyphenation applications and an Agilent 7900 ICP-MS for solution phase elemental analysis available.
Spatial Analysis of the Tissue Metabolome
Prof. Strittmatter has more than 10 years of experience in the application of imaging mass spectrometry for tissue analysis in clinical and preclinical applications – six years of which in the pharmaceutical industry. MSI can be applied to the metabolic characterization of a disease state, search and development of biomarkers and potential drug targets, and visualization of drug distribution and desired pharmacological and adverse side effects. To increase the information content, MSI can be linked to additional modalities such as hematoxylin and eosin (H&E) staining, immunohistochemical staining (IF, IHC), or imaging mass cytometry (IMC).
We predominantly develop and perform spatial metabolomics applications and studies to advance our understanding of drug delivery and drug resistance mechanisms. We work with sample types from organoids to clinical tissues. Our main instrumental platform for imaging is Desorption Electrospray Ionisation on an Orbitrap mass analyser, enabling high mass resolution and accuracy and fragmentation analysis. DESI is ideally suited for multimodal imaging workflows, enabling same section histological and immunohistochemistry-based (including highly multiplexed techniques such as imaging mass cytometry) analysis.
Metabolic Heterogeneity in Biofilms
Bacterial biofilms differ significantly in their properties from the same planktonic bacteria (cells in free suspension). For example, they exhibit significantly increased resistance to antibiotics. Our goal is to develop new methods to study the spatial heterogeneity of biofilms and establish applications in medical, biotechnological, and environmental applications. Biofilms consist of a mucus layer (extracellular polymeric matrix) in which bacteria are interconnected and are often additionally bound to an interface such as the surface of a water body or a medical implant. The extracellular polymeric substances (EPS), when combined with water, form hydrogels in which the bacteria are embedded, and nutrients and other substances are dissolved. They thus give the bacteria a stable life form and the ability to communicate with each other and exchange substances and genes.
At the microscopic level, it is known that there are gradients in nutrients, pH, and oxygen concentration. However, biofilms have been little studied for their meso- and macroscopic heterogeneity due to a lack of appropriate analytical methods. Imaging mass spectrometry offers the possibility of analyzing hundreds of microbial intra- and extracellular metabolites at the meso- and macroscopic levels for their spatial distribution. This allows biofilms to be metabolically characterized, segmented and analyzed to respond to external stressors (e.g., antibiotics, other microorganisms, and bacteriophages). Our goal is to examine and understand the biofilm as a whole organism, thus bridging the gap in understanding between microscopic and macroscopic behavior. Furthermore, such methods can help eliminate undesirable biofilms (e.g., in medicine) and improve desirable biofilms (e.g., in biotechnology).